[Feedback][Tutorial Contents][hep.lcd Home]
![]()
There are currently three possible algorithms to choose from, implemented as part of the LCD cluster finding package. They are:
All of these clustering algorithms should currently be considered as first passes which need more work. In particular there is currently no attempt made to combine clusters in different detector elements (e.g. Hadronic and EM calorimeters) and no attempt to refine clusters or match them with tracking info.
SimpleClusterBuilder![]() is a
Processor
is a
Processor![]() , so to call it you need to create a
Driver
, so to call it you need to create a
Driver![]() and add the Processor to the Driver in
the driver's constructor. (Note that Driver extends EventAnalyzer, so we can use a Driver
in JavaAnalysisStudio in the same way we used an EventAnalyzer in earlier tutorials -
except that a Driver provides its own implementation of processEvent so we do not need to
provide one ourselves.)
and add the Processor to the Driver in
the driver's constructor. (Note that Driver extends EventAnalyzer, so we can use a Driver
in JavaAnalysisStudio in the same way we used an EventAnalyzer in earlier tutorials -
except that a Driver provides its own implementation of processEvent so we do not need to
provide one ourselves.)
import hep.analysis.*; import hep.lcd.util.driver.*; import hep.lcd.recon.cluster.simple.*; final public class LCDDriver extends Driver { public LCDDriver() { add(new SimpleClusterBuilder()); add(new ClusterAnalyzer()); } }
In the above example we refer to a ClusterAnalyzer class which is used by all of the examples in this tutorial to perform a simple analysis of the Clusters created by each cluster finder. The ClusterAnalyzer class is defined below:
import hep.analysis.*; import hep.lcd.util.driver.*; import hep.lcd.event.*; import java.util.*; import hep.physics.*; final class ClusterAnalyzer extends AbstractProcessor { public final void process(LCDEvent event) { ClusterList cl = event.getEMClusters(); histogram("NClust").fill(cl.getNClusters()); Enumeration e = cl.getClusters(); while (e.hasMoreElements()) { Cluster c = (Cluster) e.nextElement(); histogram("Hits per cluster").fill(c.getNHits()); histogram("Cluster Energy").fill(c.getEnergy()); histogram("Rms R").fill(c.getRMSRadius()); histogram("R").fill(c.getEnergyRadius()); // Find the hit with the highest energy double eMax = 0; Enumeration ee = c.getHits(); while (ee.hasMoreElements()) { CalorimeterHit h = (CalorimeterHit) ee.nextElement(); double energy = h.getEnergy(); eMax = Math.max(eMax,energy); } histogram("E Max").fill(eMax); } } }
One of the advantages of using Drivers and Processors is that there a number of utility functions provided by the Driver class. One of these is the ability to set the level of diagnostic histograms generated by the Processors called by the Driver. To turn on the ClusterAnalyzers diagnostic histograms it is only necessary to add a single line to the constructor of LCDDriver in the above example.
setHistogramLevel(ProcessorContext.LEVEL_HIGH);
The SimpleClusterBuilder sets an energy threshold which a calorimeter hit must exceed before it can form the nucleus of a cluster. By default this threshold is set to 0, but it can be set using the setEMThreshold and setHadronicThreshold methods of SimpleClusterBuilder.
The ClusterCheater![]() uses the MC truth information to build
"perfect" clusters, by combining all of the hits associated with the same MC
particle into a single cluster. It is primarily useful for diagnostic purposes,
particularly for comparison with other less-perfect algorithms. The following example
shows how to call the ClusterCheater.
uses the MC truth information to build
"perfect" clusters, by combining all of the hits associated with the same MC
particle into a single cluster. It is primarily useful for diagnostic purposes,
particularly for comparison with other less-perfect algorithms. The following example
shows how to call the ClusterCheater.
import hep.analysis.*; import hep.lcd.util.driver.*; import hep.lcd.recon.cluster.cheat.*; final public class LCDDriver extends Driver { public LCDDriver() { add(new ClusterCheater()); add(new ClusterAnalyzer()); } }
The radial cluster builder gets its name from the order in which it traverses hits: from the innermost layers outward radially. It also allows individual calorimeter hits to be binned prior to clustering. The default binning is: layer-binning = 4, phi-binning = theta-binning = 2. That is, the detector is treated as if it had one quarter of the layers it actually has, and coarser phi- and theta-segmentation by a factor of two. These parameters are settable by the application independently for the EM and Had calorimeters by the methods setEMBinning and setHADBinning. This can be useful either for gauging the effect of less fine calorimeter segmentation, or for experimenting with clustering strategies. The algorithm also involves thresholds settable by the caller. For each of EM and Had there is an "ignore" threshold (all hits with energy under this value will be ignored) and a "nucleus" threshold. Any clusters all of whose hits are below this value will be excluded from the list of good clusters, but are still accessible for further analysis via methods of RadialClusterBuilder. The default value for all energy thresholds is 0.
The following example illustrates the use of the RadialClusterBuilder with no special configuration.
import hep.analysis.*; import hep.lcd.util.driver.*; import hep.lcd.recon.cluster.radial.*; final public class LCDDriver extends Driver { public LCDDriver() { add(new RadialClusterBuilder()); add(new ClusterAnalyzer()); } }
Here is an example illustrating the use of some of the RadialClusterBuilder configuration methods.
import hep.analysis.*; import hep.lcd.util.driver.*; import hep.lcd.recon.cluster.radial.*; final public class LCDDriver extends Driver { public LCDDriver() { RadialClusterBuilder clusterer = new RadialClusterBuilder(); // Double usual number of layers in HAD bins clusterer.setHADBinning(2, 2, 8); // Exclude EM clusters of only low energy hits clusterer.setEMNucleusEnergy(0.00001); add(clusterer); add(new ClusterAnalyzer()); } }
It is easy to run more than one clustering algorithm on the same event and compare the results, as the following code segment shows:
To be supplied
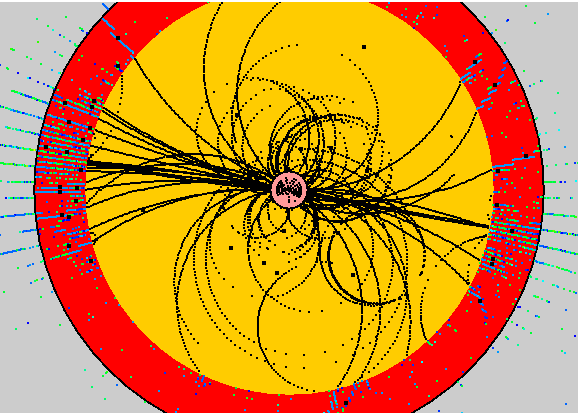
Once a clustering algorithm has been run the event display will automatically display the clusters as black diamonds. In future the cluster display will be extended to also show the rms size of the cluster.