

XML meets JASHist...
Scope
This document provides examples of how to use the XML capabilities of the JASHist package and a list of currently implemented and planned features. It also covers code level topics which interested developers will benefit from knowing. It is meant to be read from start to finish.
Prerequisites
If you want a good technical introduction to XML see this excellent, but a bit out of date, article: http://nwalsh.com/docs/articles/xml/. If you like reading specifications the annotated XML specification is here: http://www.xml.com/axml/testaxml.htm.
Terminology / Files Specific To JASHist & XML
Term
Description
plotML.dtd
The DTD (Document Type Definition) which specifies what is a legal plotML file.
mem.xml
The xml file which specifies two memory
plots. (In jas.hist.test.)
Menu
This document has two appetizers and three entrees. Appetizers are just light, tasty examples that should impress you and prepare/motivate you for the much more substantial entrees.
Appetizers
Appetizer I - A Simple 1D Histogram
Copy the following text and save it in a file called simple1DExample.xml:
<?xml version="1.0" encoding="ISO-8859-1"?> <!DOCTYPE plotML SYSTEM "plotML.dtd"> <plotML> <plot> <legend visible="true"> </legend> <dataArea> <data1d title="Hello World!"> <bins1d> 0.0,2.0,2.0 13.0,1.2,2.3 53.0,7.280109889280518,7.280109889280518 27.0,16.0,1.8 2.0,1.4142135623730951,1.4142135623730951 </bins1d> <binnedDataAxisAttributes axis="x" min="0.0" max="1.0" numberOfBins="5" type="double"/> </data1d> </dataArea> </plot> </plotML>
From the command line, ensure your CLASSPATH is properly configured. Go to the directory where you saved the file and type the following command:
java jas.hist.test.XMLHistViewer simple1DExample.xml
You should see a new Java application window that looks like this:
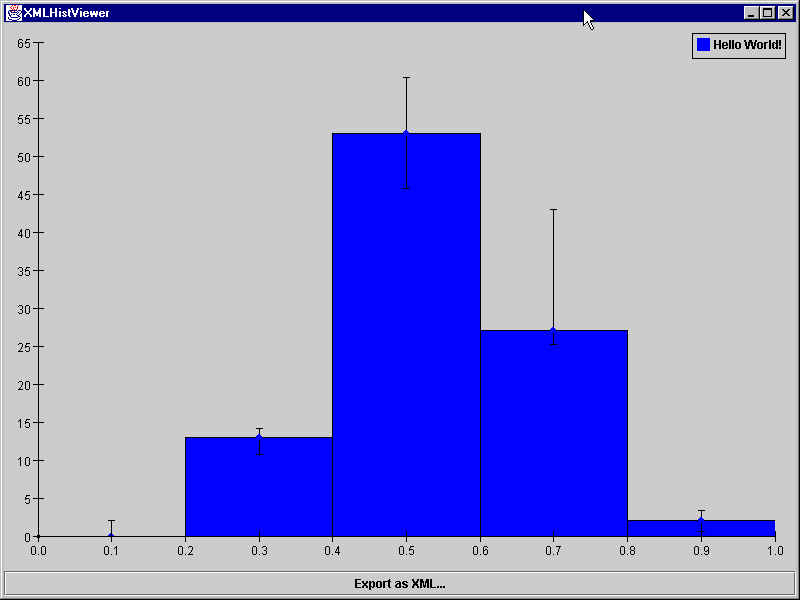
Appetizer II - A Simple 2D Histogram
Copy the following (large amount of) text and save it in a file called simple2DExample.xml:
<?xml version="1.0" encoding="ISO-8859-1" ?> <!DOCTYPE plotML SYSTEM "plotML.dtd"> <plotML> <plot> <title> <border type="Line" color="Red"/> <bounds x="5" y="5" width="395" height="42"/> <label text="2D or not 2D?"> <font face="Monospaced" points="25" style="BOLD+ITALIC"/> </label> </title> <dataArea> <border type="Bevel Out"/> <bounds x="5" y="47" width="395" height="258"/> <axis location="x0" min="0.0" max="1.0" logarithmic="false" numberOfBins="40" type="double" showOverflowBars="false"> <label text="This is the X Axis!"> <font face="SansSerif" points="22" style="BOLD"/> </label> <font face="Dialog" points="12" style="PLAIN"/> </axis> <axis location="y0" min="0.0" max="1.0" logarithmic="false" numberOfBins="40" type="double" showOverflowBars="false"> <font face="Serif" points="15" style="ITALIC"/> </axis> <data2d type="histogram2d" title="Gaussian 7" dataHistStyles="STYLE_BOX" shapeColor="Dark Gray" overFlowBinColor="Magenta" startDataColor="White" endDataColor="Black" dataOverFlow="false" dataShowPlot="true"> <bins2d xSize="40" ySize="40"> 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 4.0,2.0,2.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 1.0,1.0,1.0 5.0,2.23606797749979,2.23606797749979 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 1.0,1.0,1.0 4.0,2.0,2.0 0.0,0.0,0.0 6.0,2.449489742783178,2.449489742783178 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 1.0,1.0,1.0 4.0,2.0,2.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 5.0,2.23606797749979,2.23606797749979 3.0,1.7320508075688772,1.7320508075688772 6.0,2.449489742783178,2.449489742783178 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 1.0,1.0,1.0 5.0,2.23606797749979,2.23606797749979 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 7.0,2.6457513110645907,2.6457513110645907 4.0,2.0,2.0 6.0,2.449489742783178,2.449489742783178 5.0,2.23606797749979,2.23606797749979 5.0,2.23606797749979,2.23606797749979 1.0,1.0,1.0 4.0,2.0,2.0 0.0,0.0,0.0 5.0,2.23606797749979,2.23606797749979 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 5.0,2.23606797749979,2.23606797749979 5.0,2.23606797749979,2.23606797749979 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 4.0,2.0,2.0 7.0,2.6457513110645907,2.6457513110645907 5.0,2.23606797749979,2.23606797749979 2.0,1.4142135623730951,1.4142135623730951 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 3.0,1.7320508075688772,1.7320508075688772 4.0,2.0,2.0 8.0,2.8284271247461903,2.8284271247461903 7.0,2.6457513110645907,2.6457513110645907 6.0,2.449489742783178,2.449489742783178 4.0,2.0,2.0 8.0,2.8284271247461903,2.8284271247461903 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 5.0,2.23606797749979,2.23606797749979 6.0,2.449489742783178,2.449489742783178 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 6.0,2.449489742783178,2.449489742783178 5.0,2.23606797749979,2.23606797749979 6.0,2.449489742783178,2.449489742783178 5.0,2.23606797749979,2.23606797749979 7.0,2.6457513110645907,2.6457513110645907 8.0,2.8284271247461903,2.8284271247461903 7.0,2.6457513110645907,2.6457513110645907 7.0,2.6457513110645907,2.6457513110645907 5.0,2.23606797749979,2.23606797749979 6.0,2.449489742783178,2.449489742783178 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 4.0,2.0,2.0 7.0,2.6457513110645907,2.6457513110645907 8.0,2.8284271247461903,2.8284271247461903 4.0,2.0,2.0 7.0,2.6457513110645907,2.6457513110645907 4.0,2.0,2.0 6.0,2.449489742783178,2.449489742783178 10.0,3.1622776601683795,3.1622776601683795 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 5.0,2.23606797749979,2.23606797749979 3.0,1.7320508075688772,1.7320508075688772 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 2.0,1.4142135623730951,1.4142135623730951 6.0,2.449489742783178,2.449489742783178 7.0,2.6457513110645907,2.6457513110645907 9.0,3.0,3.0 6.0,2.449489742783178,2.449489742783178 9.0,3.0,3.0 9.0,3.0,3.0 6.0,2.449489742783178,2.449489742783178 8.0,2.8284271247461903,2.8284271247461903 6.0,2.449489742783178,2.449489742783178 2.0,1.4142135623730951,1.4142135623730951 9.0,3.0,3.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 5.0,2.23606797749979,2.23606797749979 5.0,2.23606797749979,2.23606797749979 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 7.0,2.6457513110645907,2.6457513110645907 4.0,2.0,2.0 6.0,2.449489742783178,2.449489742783178 5.0,2.23606797749979,2.23606797749979 8.0,2.8284271247461903,2.8284271247461903 9.0,3.0,3.0 7.0,2.6457513110645907,2.6457513110645907 5.0,2.23606797749979,2.23606797749979 4.0,2.0,2.0 4.0,2.0,2.0 5.0,2.23606797749979,2.23606797749979 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 3.0,1.7320508075688772,1.7320508075688772 5.0,2.23606797749979,2.23606797749979 5.0,2.23606797749979,2.23606797749979 6.0,2.449489742783178,2.449489742783178 7.0,2.6457513110645907,2.6457513110645907 5.0,2.23606797749979,2.23606797749979 7.0,2.6457513110645907,2.6457513110645907 9.0,3.0,3.0 5.0,2.23606797749979,2.23606797749979 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 3.0,1.7320508075688772,1.7320508075688772 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 1.0,1.0,1.0 4.0,2.0,2.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 7.0,2.6457513110645907,2.6457513110645907 3.0,1.7320508075688772,1.7320508075688772 5.0,2.23606797749979,2.23606797749979 5.0,2.23606797749979,2.23606797749979 1.0,1.0,1.0 10.0,3.1622776601683795,3.1622776601683795 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 3.0,1.7320508075688772,1.7320508075688772 4.0,2.0,2.0 3.0,1.7320508075688772,1.7320508075688772 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 1.0,1.0,1.0 4.0,2.0,2.0 6.0,2.449489742783178,2.449489742783178 2.0,1.4142135623730951,1.4142135623730951 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 5.0,2.23606797749979,2.23606797749979 5.0,2.23606797749979,2.23606797749979 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 4.0,2.0,2.0 1.0,1.0,1.0 4.0,2.0,2.0 3.0,1.7320508075688772,1.7320508075688772 1.0,1.0,1.0 4.0,2.0,2.0 4.0,2.0,2.0 2.0,1.4142135623730951,1.4142135623730951 3.0,1.7320508075688772,1.7320508075688772 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 4.0,2.0,2.0 1.0,1.0,1.0 7.0,2.6457513110645907,2.6457513110645907 3.0,1.7320508075688772,1.7320508075688772 6.0,2.449489742783178,2.449489742783178 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 3.0,1.7320508075688772,1.7320508075688772 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 3.0,1.7320508075688772,1.7320508075688772 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 2.0,1.4142135623730951,1.4142135623730951 4.0,2.0,2.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 3.0,1.7320508075688772,1.7320508075688772 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 2.0,1.4142135623730951,1.4142135623730951 1.0,1.0,1.0 1.0,1.0,1.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 2.0,1.4142135623730951,1.4142135623730951 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 1.0,1.0,1.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 0.0,0.0,0.0 </bins2d> <binnedDataAxisAttributes axis="x" min="0.0" max="1.0" numberOfBins="40" type="double"/> <binnedDataAxisAttributes axis="y" min="0.0" max="1.0" numberOfBins="40" type="double"/> </data2d> </dataArea> </plot> </plotML>
From the command line, ensure your CLASSPATH is properly configured. Go to the directory where you saved the file and type the following command:
java jas.hist.test.XMLHistViewer simple2DExample.xml
You should see a new Java application window that looks like this:

Entrees
There are two types of entrees: those where the plotML merely specifies the data source, and those where the plotML file is the data source.
1. The plotML File Specifies the Data Source
Entree I - Memory Plot
We will start with an example which shows how we can use a simple little XML file to specify a live data source for a plot. This example makes use of a standalone tool called XMLHistViewer which currently lives in the jas.hist.test package. In this package is a file called mem.xml which we will look at shortly; first, however, we will just run the XMLHistViewer Java application with mem.xml as its argument and see what happens.
From the command line, ensure your CLASSPATH is properly configured and type the following command:
java jas.hist.test.XMLHistViewer mem.xml
You should see a new Java application window that looks like this:
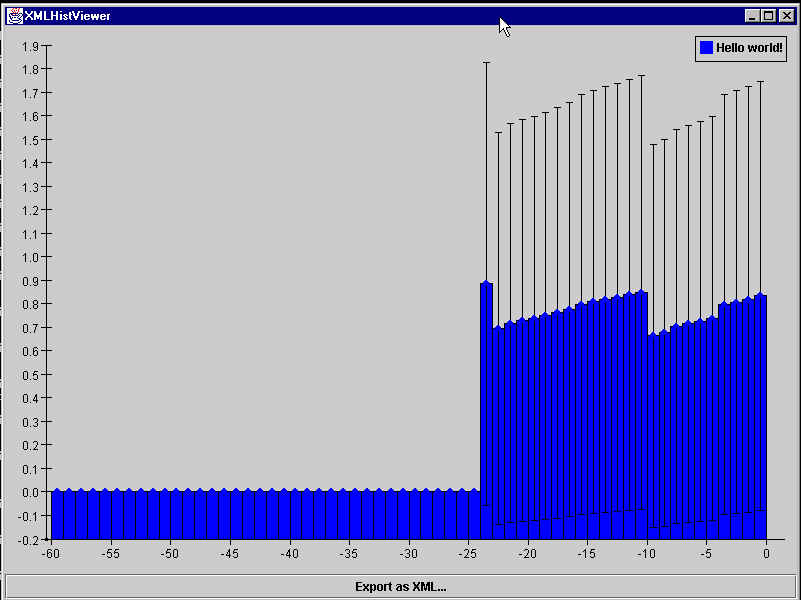
What is being displayed is the memory usage of the Java Virtual Machine. (Yes, this is the same thing that is displayed in the welcome page of Java Analysis Studio.)
Now let's look at the mem.xml file that we passed to the XMLHistViewer to get this plot:
<?xml version="1.0" encoding="ISO-8859-1" ?> <!DOCTYPE plotML SYSTEM "plotML.dtd"> <plotML> <plot> <legend visible="true"> </legend> <dataArea> <data1d> <class name="jas.hist.test.MemoryDataSource" param="Hello world!"/> </data1d> </dataArea> </plot> </plotML>
Yes, that's the whole thing. Now let's take it apart line-by-line (the lines will be in boldface monospaced font ) and see what it all means:
<?xml version="1.0" encoding="ISO-8859-1" ?>
This fairly self-explanatory line just says which version of XML we are using and the fact that the file is in ISO-8859-1 (i.e. boring old ASCII) encoding. Nothing shocking.
<!DOCTYPE plotML SYSTEM "plotML.dtd">
This is more interesting: Here we are specifying that the DOCTYPE that the mem.xml file supposedly conforms to is defined in a file called "plotML.dtd". This file is the DTD (Document Type Definition) which specifies what is a legal plotML file. The DTD provides the rules that the mem.xml file must follow.
Where is this DTD file, you ask?
In the jas.hist package.
How can we
make sure that, no matter how your system is configured, we can find this
DTD?
We're crafty, that's how! If you're
curious, take a look at the public void parse(InputStream
stream) method in jas.hist.XMLHistBuilder. The answer
involves setting the EntityResolver to one which we've subclassed and whose
resolveEntity() method we've overridden.
<plotML>
The plotML element contains plots and plotPages. It serves the same purpose as, for example, the opening and closing HTML elements do in HTML: to identify a file as conforming to the appropriate DTD. plotML is like XHTML, insofar as they are both specific "applications" of XML--XHTML for web pages, plotML for plots.
This is how the plotML element is defined in the DTD:
<!ELEMENT plotML (plot | plotPage)>
<plot>
This "opens up" a plot element. This is how the plot element is defined in the DTD:
<!ELEMENT plot ( title?, legend?, dataArea?, bounds?, border?, colorScheme? )>
In a DTD, a question mark means that a sub-element is optional. (DTDs use a lot of the standard regular expressions symbols: ? = 0 or 1, * = 0 or more, + = 1 or more, | = or.)
So this means that every element in a plot is optional. This is since the plotML designers are (as they are known to do) trying to be as flexible as possible.
<legend
visible="true">
</legend>
This opens and closes a legend element which has the visible attribute set to true. This is how the legend element is defined in the DTD:
<!ELEMENT legend (border?, bounds?, colorScheme?, font?, legendItem*)> <!ATTLIST legend visible CDATA "automatic" >
The visible attribute can have 1 of 3 values: "true", "false" and the default value of "automatic". If it is "true" then the legend is always shown; if it is "false" the legend is never shown; it it is "automatic" then the legend is only shown if the number of data sources is > 1. (This is sensible since if there is only one thing on a plot you don't need a legend.)
<dataArea>
This opens up a dataArea element. This is how the dataArea element is defined in the DTD:
<!ELEMENT dataArea ( border?, bounds?, colorScheme?, axis*, (data1d+ | data2d+)? ) >
In English, this means that a dataArea contains (in this order): border (optional), bounds (optional), colorScheme (optional), 0 or more axes, 1 or more data1d's OR 1 or more data2d's (optional). Note: The axis element contains an attribute which specifies its type. So there is no potential ambiguity in determining which axis is being specified if only one of them is. The phrase (data1d+ | data2d+)? which I have written as "1 or more data1d's OR 1 or more data2d's" means that it is legal to not specify any data1d's or data2d's (that's what the ? outside of the ()s is there for), but if you DO specify a data1d or a data2d then you must specify ALL data1ds OR ALL data2ds, i.e. you CANNOT mix and match data1d's and data2d's (since it would be rather hard to plot).
<data1d>
<class
name="jas.hist.test.MemoryDataSource"
param= "Hello
world!"/>
</data1d>
This opens up a data1d element which contains a class data source whose name is jas.hist.test.MemoryDataSource and whose parameter is "Hello world!". Then it closes the data1d element.
Now for the gory details:
This is how the data1D element is defined in the DTD:
<!ELEMENT data1d ( ( (class, (binnedDataAxisAttributes | pointDataAxisAttributes)?) | (points, pointDataAxisAttributes) | (bins1d, binnedDataAxisAttributes) ), axisLabels? ) > <!ATTLIST data1d title CDATA #IMPLIED histogramBarsFilled NMTOKEN "true" histogramBarColor CDATA #IMPLIED errorBarColor CDATA #IMPLIED dataPointColor CDATA #IMPLIED dataPointStyle NMTOKEN #IMPLIED dataPointSize CDATA "6" lineColor CDATA #IMPLIED showHistogramBars NMTOKEN "true" showErrorBars NMTOKEN "true" showDataPoints NMTOKEN "true" showLinesBetweenPoints NMTOKEN "false" > <!-- title = what is put into the legend by default histogramBarsFilled = true if the histogram bars should be filled, false otherwise histogramBarColor = a string like "Red" or "red" or "RED" -->
The definition of the the data1d element is rather convoluted since it is enforcing the following conditions:
1. A data1d contains exactly one of the following three elements: class, points, bins1d. These elements all produce data sources.
2. If a class element is specified then either the binnedDataAxisAttributes or the pointDataAxisAttributes element applies but it is optional. Howeber, if a points or bins1d element is specified then the appropriate DataAxisAttributes (point for points, binned for bins1d) element is required.
3. An axisLabels element can always be specified to override the defaults. (NOTE: It is an unresolved issue how this will work with a class element--currently we just ignore it in this case!)
That's all we need to concern ourselves with about the data1d element. We did not need to specify any of the numerous attributes (in the ATTLIST) since they all are either optional or have sensible defaults.
Now we need to take a closer look at the class element.
This is how the class element is defined in the DTD:
<!ELEMENT class EMPTY> <!ATTLIST class name CDATA #REQUIRED param CDATA #REQUIRED >
Although it is fairly small, the class element is probably the most complicated element we've examined so far. It is an empty element, which just means that it has no sub-elements. The two attributes, both of which are required, are name and param. The name attribute specifies the fully-qualified class name (in this case jas.hist.test.MemoryDataSource) of a class which MUST implement the HasDataSource interface. What is the HasDataSource interface? First of all, an interface in Java is something like a "pure" abstract class: all you can provide are method signatures (names, arguments, return types), not method bodies. (A method is a member funtion of a class.) The HasDataSource interface specifies one method: getDataSource(String param). This is the HasDataSource interface:
package jas.hist;
public interface HasDataSource {
public DataSource getDataSource(String param);
}
The getDataSource method takes a String parameter (imaginitively called param) and uses it to return a jas.hist.DataSource. What is a DataSource, you ask? Well, a DataSource is also an interface--except it is an interface which is implemented by any JASHist datasource. Specifically, it is implemented by the more detailed jas.hist.Rebinnable1DHistogramData and jas.hist.Rebinnable2DHistogramData interfaces (whose names describe them fairly well--except for the fact that they might not actually be rebinnable). And, as you might have guessed, the String which is passed in as the value of param is indeed the value of the param attribute (in this case "Hello") of the class element. That's why they're both called param.
So in a nutshell, the purpose of the class element is to specify the values of name and param. The value of param is passed as the argument to the getDataSource method of the class whose name is specified in the name attribute. If we know what we're doing, the class is guaranteed to have a getDataSource method with the signature
public DataSource getDataSource(String param);
since it implements the HasDataSource interface. The only problem that can happen is if we forget what we're doing and specify a class which does not implement the HasDataSource interface. Then all kinds of problems would happen. So parsers of plotML files (like jas.hist.XMLHistBuilder) should ensure that they check that the class which is specified in the name attribute of the class element really does implement HasDataSource.
Phew.
</dataArea>
</plot>
</plotML>
Here we are just closing off all of the elements we opened.
That's it for our first example. Go get a coffee or soda and we'll continue.
2. The plotML File IS the Data Source!
Okay, so now that we've seen how to add a dynamic data source to a plot by writing a plotML file, it's time to look at how we can specify the actual data in the plotML file itself. There are two ways to do this, since there are two kinds of data we can have: bins and points. So not surprisingly the two other kinds of sub-elements of the data1d and data2d elements are bins and points. (Yes, the data2d element can also, like the data1d element, contain a class data source--I just don't have one to show you.) We will take turns looking at examples which use the bins and the points attributes. We will arbitrarily choose to use bins with a data1d and points with a data2d.
Entree II -
The Bins Attribute with a 1D HistogramInstead of inventing a new example, let's just go through the first Appetizer code line by line. You've already seen what it looks like in the XMLHistViewer; here's the code again so you don't have to scroll up.
<?xml version="1.0" encoding="ISO-8859-1"?> <!DOCTYPE plotML SYSTEM "plotML.dtd"> <plotML> <plot> <legend visible="true"> </legend> <dataArea> <data1d title="Hello World!"> <bins1d> 0.0,2.0,2.0 13.0,1.2,2.3 53.0,7.280109889280518,7.280109889280518 27.0,16.0,1.8 2.0,1.4142135623730951,1.4142135623730951 </bins1d> <binnedDataAxisAttributes axis="x" min="0.0" max="1.0" numberOfBins="5" type="double"/> </data1d> </dataArea> </plot> </plotML>
<?xml version="1.0" encoding="ISO-8859-1"
?>
<!DOCTYPE plotML
SYSTEM "plotML.dtd">
<plotML>
<plot>
<legend
visible="true"></legend>
<dataArea>
<data1d>
The meaning of these seven lines was already explained above in the memory plot example.
<bins1d>
This line indicates that the data source being produced is a bins data source, with the bins being explicitly specified as #PCDATA.
0.0,2.0,2.0
13.0,1.2,2.3
53.0,7.280109889280518,7.280109889280518
27.0,16.0,1.8
2.0,1.4142135623730951,1.4142135623730951
These five lines are the actual bin data. Why is this just character data (which is called #PCDATA in XML) instead of, say, a bin element? This is answered in a lengthy comment in the plotML DTD, which is pasted here for your convenience:
The next three elements, bins1d, bins2d and points all have no children but
instead contain #PCDATA (which basically is unstructured character data).
This data is the bin and point data... and there is a LOT of it. A typical
2D histogram has around 2000 bins, and a typical 2D scatterplot has around
20,000 points. In theory we could make each bin or point an XML element, but
this would be a Very Bad Idea for these reasons:
1. It would waste a ton of space. Although XML is not terse, this would
be excessive even by XML standards. Visualize this repeated 2000 or
20,000 times:
<bin contents="0.0"/>
That's a lot of characters when we only need three ("0.0").
2. Many implementations (such as the one in JAS) will parse plotML files
using the DOM. Having a tree with 2000 or 20,000 nodes (one for each
bin or point) is absurd.
So we will define our own format of this data rather than use XML to do the job
for us. Yes, this means that plotML parsers have to do the extremely trivial
job of parsing this data themselves.
Here are the formats:
(Each bin or point is ALWAYS on its own line.)
A bin, whether it is part of a bins1d or a bins2d, is one comma-delimited
row of numbers. The form is: amount_in_bin, positive_error, negative_error
If positive_error = negative_error then the form is: amount_in_bin, error
If error is not specified (in which case it is assumed to be the square root),
the form is: amount_in_bin
For example:
5,2.234,1.432 the case where both error bars are specified separately
5,2.234 the case where positive_error = negative_error
5 the case where no error bars are specified
A point is always composed of comma-separated numbers. The number of these
numbers is specified by the dimensions attribute of the points element.
So points in a 2D histogram obviously have 2 dimensions.
For example:
-0.7319118445148435,-0.45307473022773015
The bin and point data SHALL NOT be tabbed in to make it look nicely formatted.
This, although easy to do, would be a very inefficient use of storage. So it
is NOT the responsibility of implementations to strip leading whitespace,
although this is trivial to do. Anyone reading a plotML file is smart enough
to understand it without the needless tabbing.
<binnedDataAxisAttributes axis="x" min="0.0" max="1.0" numberOfBins="5" type="double"/>
This specifies the various attributes of the data which are manifested on the axis. This is how the dataAxisAttributes element is defined in the DTD:
<!ELEMENT binnedDataAxisAttributes EMPTY> <!ATTLIST binnedDataAxisAttributes axis CDATA #REQUIRED min CDATA #REQUIRED max CDATA #REQUIRED numberOfBins CDATA #REQUIRED type CDATA #REQUIRED > <!-- axis = which axis does this apply to? (currently x or y) min = the smallest value on the axis max = the largest value on the axis numberOfBins = number of bins to divide the axis into (integer) type = one of "date", "string", "time", "double" -->
This is all pretty well described by the DTD comment (the <!-- ... --> thing). The DTD is fairly well-commented, so if you have a question about plotML it should be the first place you look for answers.
</data1d>
</dataArea>
</plot>
</plotML>
Here we are just closing off all of the elements we opened.
Entree III - The Points Attribute with a 2D Scatterplot
Finally, we will take a look at an example of a 2D scatterplot. Since a scatterplot plots points, we obviously cannot use a bins data source. So we get to look at our third kind of data source: the points data source.
Without further ado, let's take a look at the plotML file. Save it as scatterPlotExample.xml and view it the same way you have viewed the others. You should see the following image in the XMLHistViewer:

Here's the plotML file that produced this:
<?xml version="1.0" encoding="ISO-8859-1" ?> <!DOCTYPE plotML SYSTEM "plotML.dtd"> <plotML> <plot> <legend> <border type="Line" color="Black"/> <bounds x="297" y="5" width="103" height="58"/> </legend> <dataArea> <border type="None"/> <bounds x="5" y="5" width="395" height="308"/> <axis location="x0" min="-4.017268610886584" max="3.5369567685244125" logarithmic="false" numberOfBins="40" type="double" showOverflowBars="false"> <font face="Dialog" points="12" style="PLAIN"/> </axis> <axis location="y0" min="-8.789222050644788" max="7.430757448058924" logarithmic="false" numberOfBins="40" type="double" showOverflowBars="false"> <font face="Dialog" points="12" style="PLAIN"/> </axis> <data2d type="scatter2d" title="Hello" dataHistStyles="STYLE_BOX" shapeColor="Blue" overFlowBinColor="Red" startDataColor="White" endDataColor="Black" dataOverFlow="false" dataShowPlot="true" displayAsScatterPlot="true" dataPointSize="3" dataPointStyle="0" dataPointColor="Black"> <points dimensions="2"> -0.24906669282259855,-0.9054830906929779 -0.3016561824228778,0.6550006456594119 -1.7505464512903284,-2.34766976007338 -0.3100088905898535,0.8640600016659905 2.2844070030741372,-1.0986972214494977 -0.6062149113649464,1.3001066802854362 -1.2448344899569075,1.135593363446387 -0.1439967494510715,3.256387408657035 -0.3757464270467474,1.4904148201049559 -0.42925223224127557,-0.4744002093060208 0.006123653494437416,2.7713825489706987 2.0740475182495577,1.4284615583858338 -0.8738112522001621,-1.0187282542487919 -0.8175773227491737,1.5186421052959957 -1.8589896319782477,-1.0427612894146943 -0.49771999779804216,-4.55759644899311 0.800863119810644,0.28998088898370433 -0.6300103423296508,-3.7461046012526262 -0.2563265206052353,0.25732632951431916 0.8145123948261153,-1.2336414583599808 </points> <pointDataAxisAttributes axis="x" type="double"/> <pointDataAxisAttributes axis="y" type="double"/> </data2d> <data2d type="scatter2d" title="Whirled!" dataHistStyles="STYLE_BOX" shapeColor="Dark Gray" overFlowBinColor="Magenta" startDataColor="White" endDataColor="Black" dataOverFlow="false" dataShowPlot="true" displayAsScatterPlot="true" dataPointSize="3" dataPointStyle="1" dataPointColor="Yellow"> <points dimensions="2"> -0.42053652415579856,2.99032126521115 -0.022715418013833696,3.2025489942090863 0.10778416008382503,0.8556186021461677 0.6230357803517275,0.6113508803399765 1.5248123798028625,2.206372166895002 -0.48624781685974905,-0.735830933299497 0.5203086403916201,2.159583263689271 -1.9664912371535188,-1.7451203371386332 1.1137719717179844,0.11633507611588724 -1.756925283015025,0.7779101242508786 -1.9858328728053012,-1.9130395980242882 0.9835480894670653,-0.5549729814233773 -0.49052176993185964,4.104967298087186 -0.3145242078379201,3.2602465543142403 1.4077377151259594,-0.387250460469231 -0.95077982379975,2.138490937675827 1.2617983155776582,-0.5104170155999433 -0.11153516349610615,0.3045501225652616 1.5546708193230598,-4.011172445274846 -0.6523487663798782,-2.107087392119758 -0.41537872531974623,0.9101131873692206 -1.3779356781009757,2.652070479689068 </points> <pointDataAxisAttributes axis="x" type="double"/> <pointDataAxisAttributes axis="y" type="double"/> </data2d> </dataArea> </plot> </plotML>
Most of these elements should be familiar to you from previous examples. The key things to note in this example are:
I hope this tutorial was helpful. Please email any questions or comments to .